Progenesis QI for proteomics - What's new in the latest release?
Progenesis QI for proteomics software enables you to Quantify and then Identify the proteins that are significantly changing in your complex samples using the advantages of label-free analysis. With support for all common vendor data formats and a highly intuitive, visually guided workflow, Progenesis QI for proteomics helps you to rapidly, objectively and reliably discover proteins of interest. These can be from single or fractionated samples, using multi-group experimental designs, with the capability to handle the large sample sets typical of today's experiments. As well as conventional data-dependent analysis (DDA), Progenesis QI supports Waters MSE and HDMSE data independent analysis (DIA). Uniquely, the software also takes advantage of the additional dimension of resolution offered by Ion Mobility separations increasing peak capacity to give improvements in accuracy and precision of quantification and identification.
New Features
Improved access to peptide-level information
In v3.0 the charge deconvoluted peptide quantification results are reported, making it much easier to review and report peptide level quantification. New export options make this data readily available. Additional data drilldowns from the protein to peptide level and then to individual peptide ions have been added. The new peptide correlation scores in the tables also provide a more informed assessment of your results, as does the addition of the unique peptide column enabling tagging and filtering of proteins based on the number of unique peptides observed.
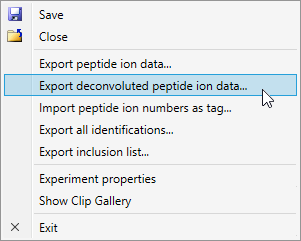
Easy export of deconvoluted peptides
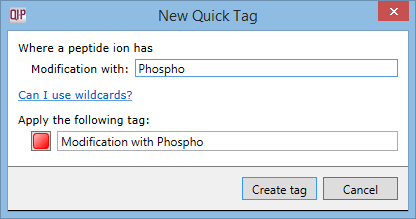
Creation of a quick tag e.g. for phospho modification:
to highlight all the peptides modified with phospho from the overall peptide list
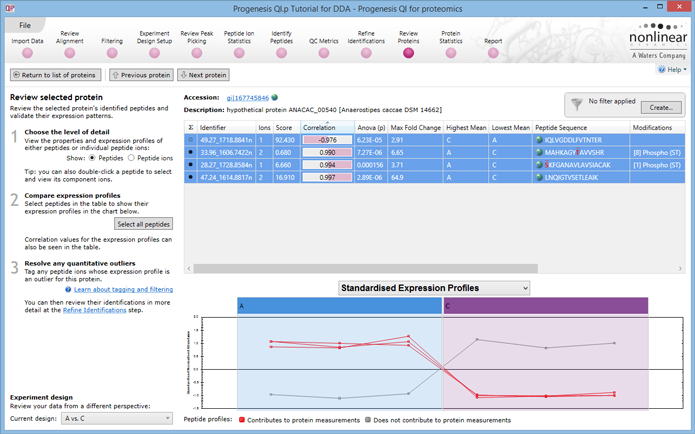
Peptide correlation at protein view helps you to see correlated peptides and enables better judgement of modifications and assignments within the data
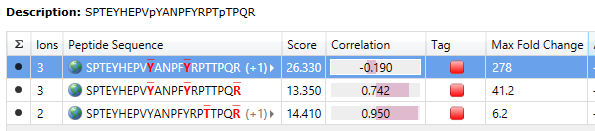
The table indicates the number of ions per peptide and the locations of any modifications
Performance improvements
Our users tell us that responsiveness and processing speed are important to them, in this version many small but significant general improvements have been made to increase responsiveness and improve areas of the workflow to give an overall improved user experience.
Improved support for data from multiple vendors
In continuity with Progenesis QI for proteomics' vendor neutral strategy, we have created a new plugin for our SCIEX users of ProteinPilot v5.0 and improved support for the generic mzML data format.





