Which runs should I use for peak picking?
When Progenesis finds the ions in your experiment, it does so by examining each of the runs selected in the Peak Picking Parameters window.
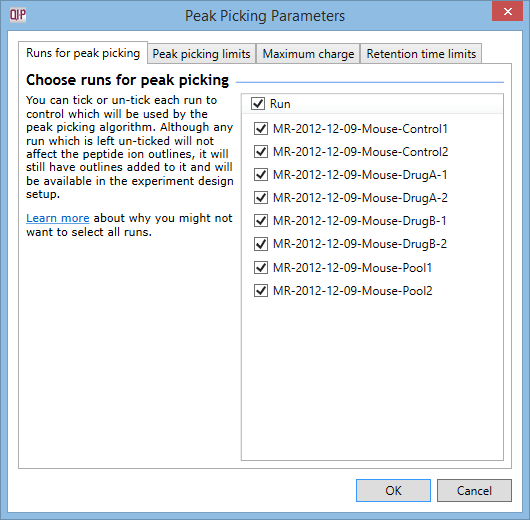
The Peak Picking Parameters window
Often, it will make sense to examine all runs during peak picking; this helps to make sure all peaks are found, even when some features are not present in all runs. Of course, it is advisable to include at least one run from each experimental condition.
There will be situations, however, in which it is appropriate to exclude some of your runs from the peak picking process. It's important to note that the resulting feature pattern will be applied to, and measured in, all runs listed in the Peak Picking Parameters dialog.
Runs not suitable for use in peak picking might include:
- In very large datasets, some runs from each test condition can be excluded to avoid over-influencing the pattern of features found
- In cross-lab studies, you may want to perform peak picking in, and later compare against, runs from only a single source e.g. a high-quality lab
- If your experiment contains a mix of MSᴱ and DDA runs, with the DDA runs sacrificing MS scans for the sake of acquiring more MSMS, you may want to exclude all DDA runs from peak picking.





