What are fragment databases and how do I use them?
Progenesis MetaScope allows you to search for compound identifications based not only on compound properties such as neutral mass, retention time and CCS, but also on the fragmentation patterns produced by compounds in your experiment.
Contents
- Fragment search methods
- What is a fragment database?
- How is the fragment database used by Progenesis MetaScope?
- How do I create my own fragment database?
- How do I append to an existing fragment database?
Fragment search methods
Progenesis MetaScope provides 3 options for how to handle fragmentation data in your experiment:
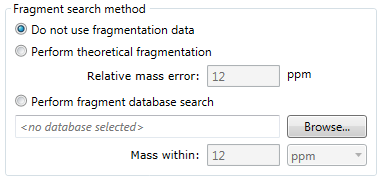
- Do not use fragmentation data
- Any fragmentation data in your experiment will be ignored.
- Perform theoretical fragmentation
- Your experimental fragmentation data is compared to theoretical fragmentation patterns generated by the simulated breaking of bonds in the structures of possible identifications.
- Perform fragment database search
- Your experimental fragmentation data is compared to known fragmentation patterns of a library of compounds, which are stored in a fragment database. The rest of this FAQ will focus on describing these fragment databases and how you can create your own.
What is a fragment database?
A fragment database is a simple text-based file (in the NIST MSP file format [pdf])
Each database file contains several compounds, and for each compound it contains some general information (ID, name, formula, adduct form), and the compound's fragmentation pattern as a series of [m/z, intensity] pairs.
It does not contain information such as neutral mass, retention time, CCS, or compound structure.
For this reason, one must combine a fragment database with a traditional SDF or CSV file containing these properties (Why is fragmentation data required to be in a separate file?).
To use a fragment database for a search, choose the SDF or CSV file containing compound properties in the Compound database section, AND an MSP fragment database file in the Fragment search method section, e.g.
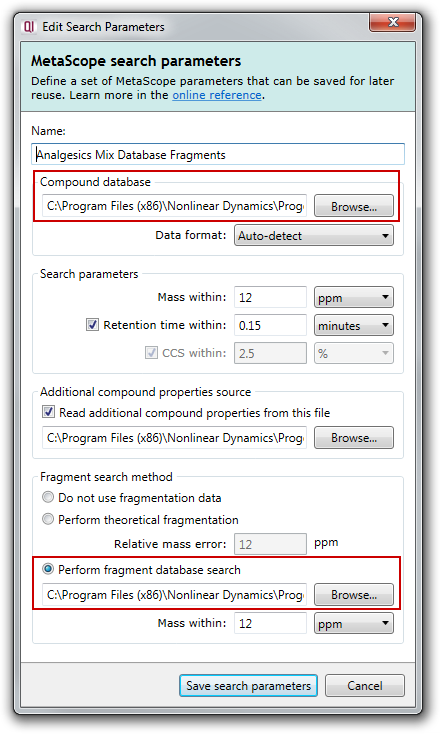
How is the fragment database used by Progenesis MetaScope?
Progenesis MetaScope will first find all possible identifications based on neutral mass, retention time, and CCS (although retention time and CCS search parameters are optional).
For each of these possible identifications, Progenesis MetaScope will then search the fragment database for a record with a matching compound ID. If a record is found, it will compare the data therein to your experimental fragmentation data, producing a "mirror plot":
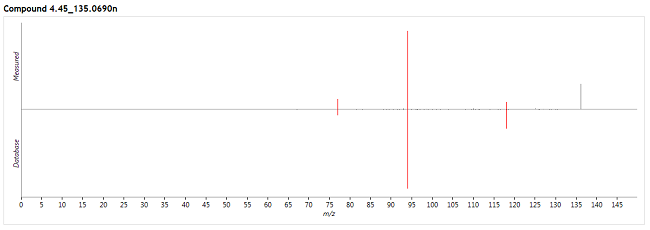
The red peaks are where a match between the experimental and database fragments was found, and the grey peaks show those with no match. Above we can see that we measured a fragment peak at around 137 Da, but this had no corresponding match in the fragment database.
After performing a search with a fragment database, Progenesis QI also provides you with a Fragmentation score, which is a metric describing how well your measured fragmentation matched that of the fragment database, e.g.
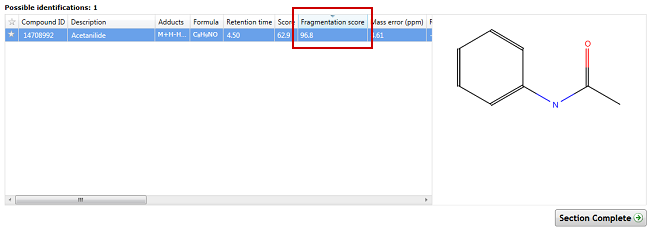
Above we can see that this match produced a very good score of 96.8 (scores range from 0-100).
How do I create my own fragment database?
Progenesis QI allows you to create your own fragment databases from your measured fragmentation patterns, which can be used for searches in future experiments.
To do this, follow the normal identification workflow (using either no fragmentation or theoretical fragmentation).
Once you have accepted your compound identifications, choose the Export fragment database... item in the File menu:
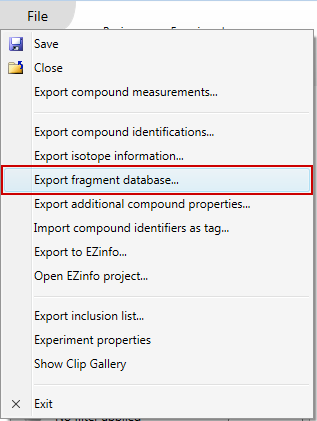
This will bring up a dialog showing you all of your accepted compound identifications. You can choose to export all or a subset of these by using the checkboxes in the first column:
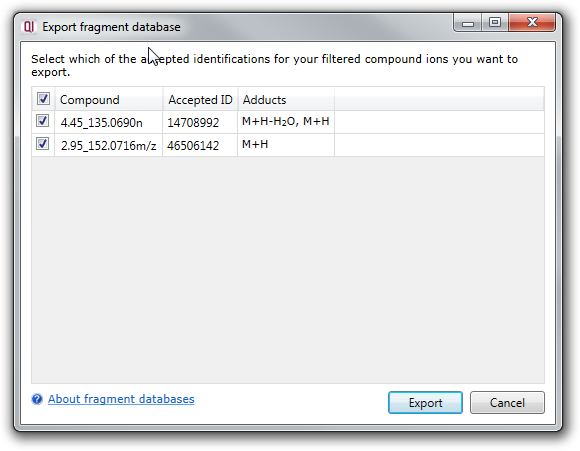
Clicking the Export button will prompt you to choose a filename for your fragment database, then carry out the export for you.
You can now use the generated fragment database in future experiments. Simply browse to the generated MSP file in the Perform fragment database search section of the MetaScope search parameters dialog (see above).
How do I append to an existing fragment database?
Suppose you have already generated your fragment database using the above method, but you wish to extend it by adding new fragmentation data from a recent experiment.
This is easy to do in Progenesis QI. When exporting the fragment database, choose the existing fragment database file when prompted for a filename.
Progenesis QI will then ask you if you wish to overwrite or append to the existing file:
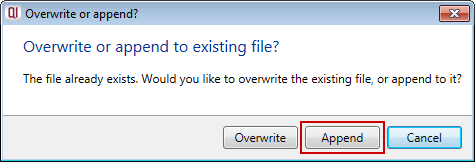
To append your new fragmentation information, choose Append, and the new data will be added to the end of your fragment database file.
Warning: Choosing Overwrite in the above dialog will completely overwrite the existing database. Make sure this is what you intend to do before choosing this option, and we recommend making a backup of the file just in case you need it again.






